INTRODUCTION
This chapter describes virology test methods applicable to the development of biological product drugs, such as recombinant proteins, subunit vaccines, therapeutic monoclonal antibodies, and growth hormones. Several topics are excluded from the scope of this chapter:
- Blood- and plasma-derived products as well as whole blood and plasma products used directly in transplantation or infusion. However, the basic principles, strategies, and testing methods for ensuring virus-free products are applicable.
- Methodologies for the safety testing of live viral vaccines.
-
Specific methods for viral clearance studies, which are described in the USP general information chapter Viral Safety Evaluation of Biotechnology Products Derived from Cell Lines of Human or Animal Origin
 1050
1050 .
.
Virology test methods have historically been employed in the clinical settings of disease diagnosis, intervention, and containment; but the development of biological (biologics and biotechnology-derived) products and therapies for human or animal use has created the need for sensitive viral detection assays for use in the GMP production and testing of biological products. This need is not limited to the production of viral vaccines, but also applies to the development and manufacture of recombinant proteins, cell and gene therapies, and other products.
Sensitive virology test methods for quality control of biological products are necessary for several reasons. The production of biological products often requires a variety of raw materials and processing reagents of animal origin that have varying potential for introducing viral contaminants. The production of biological products may allow the replication of adventitious agents during processing, and therefore these materials must be prescreened to avoid the opportunity for contamination of the product. Another point to consider regarding screening these materials is that the product may not be compatible with processing methods used to eliminate or inactivate these adventitious agents. Because of the nature of the biological products, the production process needs to include appropriate testing regimens that monitor the possible introduction of adventitious agents and/or viral agents into the systems used. For these reasons, sensitive viral detection methods are required not only for the release testing of biological drug products, but also during the intermediate stages of processing, process development, and routine manufacture. Important stages for consideration include the development of cell substrates and banks, raw materials of animal origin, process intermediates, and critical excipients when derived from animal tissues. This strategy should be augmented with viral clearance and inactivation studies whenever possible.
For products intended to contain live viruses (e.g., infectious oncolytic viruses and live viral vector products used for gene therapy), the cell- and animal-based infectivity methods discussed in this chapter may be useful only following neutralization of the specific viral entity contained in the product for any product that is intended to contain live viruses (e.g., infectious oncolytic viruses and live viral vector products used for gene therapy). Alternatively, selection of appropriate indicator cell lines or animal models in which the specific viral entity is known not to replicate can be considered. It should also be expected that assay systems based on detection of viral particles or viral components will indicate the presence of the viral entity itself in such products, but may not indicate the viability of the virus. The remainder of the chapter is divided into three sections discussing assays for the three topics: (1) Detection of Viable Viruses, (2) Detection of Viral Components, and (3) Detection of Antibodies to Viral Antigens. The chapter covers the classic virology methods that are still routinely used, as well as modern molecular and immunological approaches. The methods described in these sections may possess different sensitivities to diverse viruses; they are therefore intended to complement each other to provide a science-based foundation for the detection of adventitious viruses. Multiple methods may be used in complementary fashion to improve the pathogen safety margin of a product. Identification of viruses detected in cell-based assays on the basis of cytopathic effects often depends on the use of molecular and immunological analyses; these analyses are therefore relevant both to viral detection and to subsequent viral identification. The chapter provides an overview of the detection and analysis of the most important groups of viruses as well as the most commonly used techniques. Tests specific to individual vaccines or biological products are excluded, because they are expected to be included in monographs for such products.
Methods that are well established with little variation in practice are described in more detail, whereas methods that are more flexible are described in general terms, both in the performance of the tests and in considerations for acceptance. Relevant regulatory references are given in the Appendix. Relevant USP general chapters should be consulted with regard to bioassay design, data analysis, interpretation, and assay validation.
DETECTION OF VIABLE VIRUSES
Infectious virus particles contaminating biologics and biotechnology-derived products are of great safety concern, because they have the potential for causing serious, possibly life-threatening, infections in the patients treated. This is particularly true if the patients are immunocompromised. Although complete assurance of viral safety for finished biological products can never be realized, a significant safety margin can be established through viral detection methods applied to unprocessed bulk and raw materials before purification in combination with purification processes that demonstrate the ability to inactivate or remove potential viral contaminants present at levels too low to detect. (See Viral Safety Evaluation of Biotechnology Products Derived from Cell Lines of Human or Animal Origin  1050
1050 for information on viral clearance and inactivation methods.) For cell and gene therapy products that lack extensive purification steps, final products may be directly tested for the presence of relevant contaminating viruses. This section describes two broad systems for the detection of infectious virus: cell culture–based infectivity assays and in vivo infectivity assays. These systems may possess complementary sensitivities for viruses, and as a result, both methods may be used as limit tests for cell bank and raw material characterization and for lot release testing of biologics and biotechnology-derived products. Considerations for optimizing sample preparation for these tests are discussed, followed by a description of the more commonly employed detection assays. Finally, to ensure the reliability of experimental results, quality control issues in general and detection limit estimation in particular are discussed.
for information on viral clearance and inactivation methods.) For cell and gene therapy products that lack extensive purification steps, final products may be directly tested for the presence of relevant contaminating viruses. This section describes two broad systems for the detection of infectious virus: cell culture–based infectivity assays and in vivo infectivity assays. These systems may possess complementary sensitivities for viruses, and as a result, both methods may be used as limit tests for cell bank and raw material characterization and for lot release testing of biologics and biotechnology-derived products. Considerations for optimizing sample preparation for these tests are discussed, followed by a description of the more commonly employed detection assays. Finally, to ensure the reliability of experimental results, quality control issues in general and detection limit estimation in particular are discussed.
Sample Selection of and Preparation for Cell- and Animal-Based Virus Detection Assays
The requirements for selection, preparation, and storage of test samples for viral detection methods (cell- and animal-based) are dictated by the lability of the viruses being detected. The ability of a virus to remain infectious in the absence of a host cell is highly variable. Virus infectivity also may differ in sensitivity to repeated freezing and thawing cycles.
Sample preparation typically involves storage of test samples at low temperatures (ideally 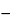 60
60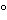 or below) as soon as practicable upon collection. When intended for use in a viral screening assay, aliquots of samples should be prepared to avoid multiple freezing and thawing. Samples intended for viral infectivity assays are typically shipped with sufficient dry ice to last several days more than the expected time required for transit. When received at the testing laboratory, the sample should be examined to verify that it is still frozen, and appropriate documentation should be completed. For any storage or hold condition, the impact of the condition on viral viability should be empirically assessed and sufficient cold chain management ensured.
or below) as soon as practicable upon collection. When intended for use in a viral screening assay, aliquots of samples should be prepared to avoid multiple freezing and thawing. Samples intended for viral infectivity assays are typically shipped with sufficient dry ice to last several days more than the expected time required for transit. When received at the testing laboratory, the sample should be examined to verify that it is still frozen, and appropriate documentation should be completed. For any storage or hold condition, the impact of the condition on viral viability should be empirically assessed and sufficient cold chain management ensured.
Typical sample types for viral detection assays are described below.
Cell Lysates—
Test samples derived from cell substrates (master and working cell banks, end-of-production cell samples) are prepared in a manner that allows sampling of both the cells (for cell-associated viruses) and the conditioned medium (for virus shed into the medium). To achieve this, a culture of the cells is sampled. A cell suspension of ~107 cells per mL in conditioned medium is prepared and frozen (ideally at 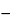 60
60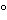 or below). Because this medium does not contain cryopreservative, the majority of the cells will lyse upon thawing of the sample, releasing the cell-associated virus. Low-speed centrifugation will remove larger cellular debris and yield a supernatant that may be inoculated directly onto detector cells in cell-based viral infectivity assays. A similar sample is prepared for in vivo viral adventitious agent testing. In this case, however, the test sample is thawed and injected without clarification into the various animal systems via the various described routes.
or below). Because this medium does not contain cryopreservative, the majority of the cells will lyse upon thawing of the sample, releasing the cell-associated virus. Low-speed centrifugation will remove larger cellular debris and yield a supernatant that may be inoculated directly onto detector cells in cell-based viral infectivity assays. A similar sample is prepared for in vivo viral adventitious agent testing. In this case, however, the test sample is thawed and injected without clarification into the various animal systems via the various described routes.
Biotechnology Bulk Harvest (Unprocessed Bulk Harvest) Samples—
Routine lot testing of bulk harvest samples is mandatory for most types of biologics (see Viral Safety Evaluation of Biotechnology Products Derived from Cell Lines of Human or Animal Origin  1050
1050 ). The sampling must be done at the unprocessed bulk harvest stage, because downstream purification processes may remove or inactivate any viruses that might contaminate the starting materials. The harvest sample from the bioreactor should be collected and stored without further manipulation, as soon as practicable, at
). The sampling must be done at the unprocessed bulk harvest stage, because downstream purification processes may remove or inactivate any viruses that might contaminate the starting materials. The harvest sample from the bioreactor should be collected and stored without further manipulation, as soon as practicable, at 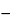 60
60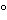 or below. To prevent multiple freeze and thaw cycles, individual aliquots should be prepared for each individual assay to be performed. Additional aliquots should be retained in case repeat testing is required. Depending on the nature of the manufacturing process, the bulk harvest samples may contain varying quantities of the production substrate cells. Because the bulk harvest does not contain cryopreservatives, the majority of the cells present will lyse upon thawing of the sample, releasing the cell-associated virus. Low-speed centrifugation to clarify the sample will result in a supernatant that may be inoculated directly onto detector cells in cell-based viral infectivity assays. There may be instances where the test sample is cytotoxic to the detector cells of the cell-based assays and procedural modifications may be required to deal with this. A similar sample is prepared for in vivo viral safety testing. In this case, however, the test sample is thawed and injected without clarification into the various animal systems via the various described routes.
or below. To prevent multiple freeze and thaw cycles, individual aliquots should be prepared for each individual assay to be performed. Additional aliquots should be retained in case repeat testing is required. Depending on the nature of the manufacturing process, the bulk harvest samples may contain varying quantities of the production substrate cells. Because the bulk harvest does not contain cryopreservatives, the majority of the cells present will lyse upon thawing of the sample, releasing the cell-associated virus. Low-speed centrifugation to clarify the sample will result in a supernatant that may be inoculated directly onto detector cells in cell-based viral infectivity assays. There may be instances where the test sample is cytotoxic to the detector cells of the cell-based assays and procedural modifications may be required to deal with this. A similar sample is prepared for in vivo viral safety testing. In this case, however, the test sample is thawed and injected without clarification into the various animal systems via the various described routes.
Raw Materials of Animal Origin—
Ingredients of animal origin used in the manufacture of biological products for human or veterinary use must be tested for species-specific viruses of concern as described in 9 CFR 113.53 (see also the USP general information chapter Bovine Serum  1024
1024 , being prepared for future publication). The raw materials may be stored under a variety of conditions, as appropriate to the raw material. Sample preparation and method of application to the test system depend on the nature of the sample. The possibility that animal-derived raw materials may contain bacterial or fungal contaminants should be considered. In some cases, it may be necessary to treat the samples with antibiotics or to filter the samples (0.22 or 0.45 micron pore size) prior to inoculation in order to prevent bacterial or fungal outgrowth in the test system. Animal sera are typically received frozen and are thawed and incorporated into the growth medium at an appropriate concentration (typically 15%, v/v) as a means of exposing the detector cells. Powdered trypsin (not less than 5 grams, as per 9 CFR 113.53) is suspended in a suitable diluent, such as phosphate-buffered saline, and is then subjected to high-speed centrifugation to pellet any virions that may be present. The concentrated pellet is resuspended in phosphate-buffered saline, and the resulting material is used to inoculate appropriate detector cells. Medium additives, such as bovine thrombin, may be incorporated into the growth medium at a predetermined multiple of the nominal concentration to be used in the manufacturing process. The resulting growth medium containing the additives is then used as a means of exposing the detector cells to the test material. The exact multiple to be used in such testing may be limited by such factors as solubility in growth medium or cytotoxicity to the detector cells. These factors should be assessed in advance of testing. The principle of using higher concentrations in the detection method than during processing should be followed, within the bounds of indicator cell toxicity, as a means to increase sensitivity to detection.
, being prepared for future publication). The raw materials may be stored under a variety of conditions, as appropriate to the raw material. Sample preparation and method of application to the test system depend on the nature of the sample. The possibility that animal-derived raw materials may contain bacterial or fungal contaminants should be considered. In some cases, it may be necessary to treat the samples with antibiotics or to filter the samples (0.22 or 0.45 micron pore size) prior to inoculation in order to prevent bacterial or fungal outgrowth in the test system. Animal sera are typically received frozen and are thawed and incorporated into the growth medium at an appropriate concentration (typically 15%, v/v) as a means of exposing the detector cells. Powdered trypsin (not less than 5 grams, as per 9 CFR 113.53) is suspended in a suitable diluent, such as phosphate-buffered saline, and is then subjected to high-speed centrifugation to pellet any virions that may be present. The concentrated pellet is resuspended in phosphate-buffered saline, and the resulting material is used to inoculate appropriate detector cells. Medium additives, such as bovine thrombin, may be incorporated into the growth medium at a predetermined multiple of the nominal concentration to be used in the manufacturing process. The resulting growth medium containing the additives is then used as a means of exposing the detector cells to the test material. The exact multiple to be used in such testing may be limited by such factors as solubility in growth medium or cytotoxicity to the detector cells. These factors should be assessed in advance of testing. The principle of using higher concentrations in the detection method than during processing should be followed, within the bounds of indicator cell toxicity, as a means to increase sensitivity to detection.
Whole Cells—
Intact viable cells are used as the test sample in certain viral detection assays. Because the test cells may attach and proliferate in the culture vessel along with the detector cells, assays using this type of sample are referred to as cocultivation assays. The requirements for the specific assay may vary in relative proportions of detector and test cells, viability of the test cells, or the confluency of test cells at the time of collection.
Cell Culture-Based Viral Detection Methods
To ensure the absence of adventitious viral agents, cell culture–based viral detection assays are used for a variety of purposes, including but not limited to clinical diagnostic procedures; evaluation of raw materials and cell substrates; assessments of the viral identity, the purity, and the potency of virus seed stocks; and lot release testing of unprocessed bulk harvests during biologics production. An important distinction between cell-based assays and direct detection assays (see the section Detection of Viral Components) is that the former will detect only replicating virus, whereas the latter will detect viral antigens, viral genomic material, and the like, which may or may not be indicative of the presence of replicating virus. Similarly, detection of circulating antibodies directed against viral antigens (discussed in the section Detection of Antibodies to Viral Antigens), may be indicative of either a current or a past infection of an animal and does not necessarily indicate that the animal is currently harboring an infection.
Infectious viruses detected in cells or in cell-derived materials fall into two broad categories, based on the expectations of the analyst. Endogenous viruses are those normally detected in the cells as a result of the integration of the viral genomic material into the host cell DNA. Exogenous viruses are those not normally present in the cells but found as a result of a viral infection of the cells.
The underlying assumption for all cell-based viral detection methods is the ability of viruses to replicate in an appropriate host cell. Viruses lack the cellular machinery required for producing their own genomic material and structural proteins, and they must therefore enter and subordinate a host cell for this purpose. Cell-based viral infectivity assays use indicator (detector) cells that serve as host cells for viable virions present in test samples.
Cell-based infectivity assays may be placed in three broad categories on the basis of types of viruses to be detected: (1) retroviral assays, (2) virus-specific assays, and (3) viral screening assays. The types of endpoints used to detect the viruses may differ by category. Although screening assays are typically not optimized for single viral entities, the virus-specific assays and titration assays, as well as some of the retroviral assays, may be optimized to some extent for specific viruses. Accurate titration of stock viruses that are used as positive controls or are used to determine the detection limit of an assay is critical.
The regulatory guidance underlying the various viral safety tests depends on the nature of the samples to be evaluated, and analysts are referred for more detail to documentation relevant to their own regulatory environments.
General Requirements for Cell Culture–Based Assays
Detector Cells and the Concept of Viral Host Range—
The range of viruses detectable using a cell-based infectivity assay depends on a number of factors, including the type of host cell(s) used as the indicator (detector) cultures and the detection endpoints used in the assay. Viruses differ in their abilities to infect specific host cell types. Most viruses exhibit at least some degree of host cell tropism (i.e., ability to infect a specific species or tissue type). This attribute is typically due to a requirement for interaction of a virion with a specific cell membrane receptor during the process of infection of the host cell. A cell susceptible to infection and capable of production of progeny by a given virus is referred to as permissive for that virus; cells not supporting viral proliferation are referred to as nonpermissive, or restricted, for that virus. As a consequence of the differences in host cell tropism, assays intended to screen for a wide range of viruses must include multiple detector cell types. For the same reasons, design of a cell-based infectivity assay for a specific virus must include a detector cell known to be permissive for that virus.
Virus Susceptibility of Common Cell Lines—
For most endpoint assays used to determine whether a host cell is infected with a virus, a monolayer culture is preferable to a semiadherent or suspension culture. For instance, cytopathic effect and hemadsorption are visualized microscopically. Cells that are not adherent have little morphology to evaluate, and hemadsorption cannot be properly evaluated in a suspension culture. For this reason, some regulatory documents pertaining to cell-based virus infectivity assays stipulate the use of monolayer detector cultures.
A list of commonly employed indicator cell lines and their application in viral screening assays is provided in Table 1. Regarding the viral tropism of these cells, “Points to Consider in the Characterization of Cell Lines Used to Produce Biologicals” (1993) and ICH's Q5A (R1) guidelines (for these references, see Appendix) require that human diploid cells such as MRC-5 and WI-38, which are permissive for a range of viruses of human concern, and monolayer cultures of the same species as that of the cell substrate used to produce the product are included in the viral screening test for biologics destined for use in humans.
Table 1. Indicator (Detector) Cell Lines Used for Adventitious Viral Screening Assays
| Cell Linea | Origin | Endpoint(s)b | Target virus(es) |
|---|---|---|---|
| Cell lines with relatively broad viral tropism: | |||
| BHK-21 | Syrian hamster | CPE, HAd, HA | Insect-borne viruses (arboviruses) |
| Vero | African green monkey | CPE, HAd, HA | Viruses infectious to humans, primates |
| For processes involving human cell substrates: | |||
| HeLa | Human | CPE, HAd, HA | Viruses infectious to humans |
| MRC-5 | Human | CPE, HAd, HA | Viruses infectious to humans |
| For processes involving Chinese hamster cell substrates: | |||
| CHO-Kl | Chinese hamster | CPE, HAd, HA | Viruses infectious to Chinese hamsters |
| For processes involving mouse cell substrates: | |||
| MEF | Mouse | CPE, HAd, HA | Viruses infectious to mouse cells |
| NIH/3T3 | Mouse | CPE, HAd, HA | Viruses infectious to mouse cells |
| For processes involving bovine cell substrates or bovine raw materials:c | |||
| MDBK | Bovine | CPE, HAd, HA | Bovine viruses |
| BT | Bovine | CPE, HAd, HA | Bovine viruses |
| EBTr | Bovine | CPE, HAd, HA | Bovine viruses |
|
a
Examples of cell lines used for viral screening assays are shown. MRC-5 and Vero, or cells with similar host ranges, are used in all assays. Depending on the cell substrate used to manufacture a biologic, additional cell lines are also used in the screening assay. In addition, a bovine cell might be included if bovine serum was used in the manufacturing process.
b
CPE, cytopathic effect; HAd, hemadsorption; HA, hemagglutination (optional).
c
Inclusion of a bovine cell in a virus screen should not be construed as a replacement for or alternative to a raw materials test. Raw materials testing is driven in the U.S. by 9 CFR 113.47 and 113.52, and cell lines used for this testing are described in Table 2.
|
|||
A list of commonly employed indicator cell lines and their application in raw materials testing assays is provided in Table 2.
Table 2. Indicator (Detector) Cell Lines Used in Raw Material Testing
| Cell Linea | Assay Type | Endpoint(s)b | Animal Origin of Raw Material |
|---|---|---|---|
| Vero | Isolation/detection | CPE, HAd, IFA | All sources |
| BT | Isolation/detection | CPE, HAd, IFA | Bovine; all sources (BVDV)c |
| EBTr | Isolation/detection | CPE, HAd, IFA | Bovine; all sources (BVDV)c |
| MDBK | Isolation/detection | CPE, HAd, IFA | Bovine; all sources (BVDV)c |
| PT-1 | Isolation/detection | CPE, HAd, IFA | Porcine |
| PK-1 | Isolation/detection | CPE, HAd, IFA | Porcine |
| MDCK | Isolation/detection | CPE, HAd, IFA | Canine |
| GT | Isolation/detection | CPE, HAd, IFA | Caprine |
|
a
The requirement (9 CFR 113.47 and 113.52) for evaluating raw materials of animal origin is to use (1) Vero cells, (2) a bovine cell for detecting BVDV, and (3) a cell line of the same species of origin as the raw material for detecting viruses of concern from that species. Examples are given of some cell lines that are used in the industry.
b
CPE, cytopathic effect; HAd, hemadsorption; IFA, immunofluorescent antibody staining.
c
As per 9 CFR 113.47, raw materials of any animal origin are to be tested for bovine viral diarrhea virus (BVDV).
|
|||
There may be very specific requirements for detector cells for certain viruses. For instance, assays intended to detect infectious HIV use human peripheral blood lymphocytes and involve a p24 antigen capture enzyme immunoassay endpoint. A list of commonly employed indicator cell lines and their application in the detection of specific viruses is provided in Table 3.
Table 3. Indicator (Detector) Cell Lines Used for Detection of Specific Virus(es)
| Cell Linea | Assay Type | Endpoint(s)b | Target Virus |
|---|---|---|---|
| 324K | Isolation/detection | CPE, HAd, IFA | Murine minute virus |
| A9 | Isolation/detection | CPE, HAd, IFA | Murine minute virus |
| BHK-21 | Isolation/detection | CPE, HAd | Arbovirusesc |
| MRC-5d | Isolation/detection | CPE | Human cytomegalovirus |
|
a
Examples of cell lines used for optimizing the detection of specific viruses or virus types are shown. In many cases, the assay methodologies must also be optimized for detection of the target viruses.
b
CPE, cytopathic effect; HAd, hemadsorption; IFA, immunofluorescent antibody staining.
c
Insect-borne viruses as a group are referred to as arboviruses. This term has no taxonomic significance.
d
Other human diploid cell lines such as WI-38 are also suitable. Assay duration must be 28 days at a minimum.
|
|||
Growth Requirements for Detector Cells—
Viral proliferation within a permissive host cell may be dependent on the rate of host cell proliferation. This is especially true for viruses that display cell-cycle dependence for generation of viral progeny. For most detection assays, detector cultures are seeded at a density intended to achieve a cell monolayer in exponential growth. This corresponds to a cell confluency of 50% or less (optimal cell densities may depend on the assay type and the detector cell to be used) at the time of inoculation of the cultures with virus or test sample. For the same reasons, the assay design may include provision for detector cell subculture (the collection of cells from the original culture and seeding of a predetermined fraction of these into a new flask). Alternatively, a passage may be performed, consisting of collection of conditioned medium from the original culture and inoculation of this material onto a secondary detector cell culture that is in log-phase growth. The frequency of subculture/passage required in an assay is determined largely by the rate of growth of the detector cell. Incorporation of these steps into detection assay designs helps to ensure that the conditions remain optimal for amplification of viral progeny within the host cells.
Need for Detector Cell Identification and Banking—
Detector cells used for cell-based viral detection assays are a critical reagent for ensuring viral safety, viral potency, viral identity, and viral clearance capacity in purification schemes. In many cases this testing is intended to support GMP processes; therefore, the detector cell banks may need to be prepared and qualified in much the same manner as other critical reagents. The details of the viral safety evaluation methods are described in Viral Safety Evaluation of Biotechnology Products Derived from Cell Lines of Human or Animal Origin  1050
1050 . Regardless of specific compliance requirements, periodic identification and qualification of the detector cell banks to be used subsequently for viral infectivity assays is good practice. The following should be regarded as minimal quality control testing for such detector cell banks: sterility, mycoplasma and viral screening, and cell identity by DNA fingerprinting, karyology, or isoenzyme analysis. In addition, specific assays for bovine and porcine viruses may be required if bovine or porcine raw materials were used in preparation of the banks. Bovine Serum
. Regardless of specific compliance requirements, periodic identification and qualification of the detector cell banks to be used subsequently for viral infectivity assays is good practice. The following should be regarded as minimal quality control testing for such detector cell banks: sterility, mycoplasma and viral screening, and cell identity by DNA fingerprinting, karyology, or isoenzyme analysis. In addition, specific assays for bovine and porcine viruses may be required if bovine or porcine raw materials were used in preparation of the banks. Bovine Serum  1024
1024 and relevant serum-type specific ancillary materials monographs should be consulted when serum products are used in cell growth media.
and relevant serum-type specific ancillary materials monographs should be consulted when serum products are used in cell growth media.
Endpoints for Detection of Viral Infection
The various endpoints used to identify infection of a detector cell with a virus include the following:
- Visual observation of cytopathic effects
- Hemagglutination or hemadsorption of erythrocytes
- Immunofluorescent staining
- Cocultivation with other types of detector cells
- Quantitative polymerase chain reaction (qPCR) for direct detection of viral genomic sequences
- Electron microscopic analysis of viral pellets or fixed cells for visual observation of viral particles
- Biochemical endpoints such as reverse transcriptase assays, which detect virus-specific enzymatic activity
These various endpoints are used in a complementary fashion, because a given virus may not cause a positive response in each endpoint. For instance, some viruses can grow to high titers without producing visible cytopathic effects and so must be detected using other endpoints. Polymerase chain reaction (PCR) and electron microscopic analysis per se are not capable of distinguishing viable from nonviable viruses. However, when used in conjunction with cell culture growth kinetics, these approaches can be powerful orthogonal detection methods to demonstrate the increase of viral replication and therefore viable virus. The failure to observe viral particles in electron microscopic analysis of fixed cells should not be considered absolute proof of the absence of infectious virus in the cells. In a general sense, the same is true for each of the detection endpoints discussed above. Each endpoint has a detection limit below which a virus may be present but not detected.
Viral Cytopathic Effects—
Visually observable manifestations of the infection of susceptible host cells with certain types of viruses are collectively referred to as viral cytopathic effects (CPE). Although CPE may be considered an indirect detection of viral infection, in the context of specific host cells they can have distinctive morphological manifestations. These may include the appearance of inclusion bodies, abnormal cell morphology, changes in culture confluency, cell death and cell lysis, and others. The nature of the CPE observed may depend on the host cell and the infecting virus. In addition, for a virus that normally causes CPE, there may exist variants that do not cause CPE. CPE can be differentiated from cytotoxic effect by the tendency of the former to exhibit progression irreversibly with time, whereas the latter may be reversible. In some cases structural proteins of viruses may cause cytotoxic effects similar to the cytopathic effects of the infectious virus. Differentiating the cytotoxic effect of such proteins from the cytopathic effect of infectious virus may require observation of the culture over time to determine whether the effect progresses or the cells appear to recover. Alternatively, cell-free passage of the original culture onto fresh detector cells can be used to differentiate these two apparently similar manifestations. The cytotoxicity associated with the structural proteins in the absence of infectious virus would not be expected to pass to the secondary culture.
Giemsa staining may optimize the ability of the operators to visualize certain inclusion bodies (clusters of viral particles) that are characteristic of viral cytopathic effect and is required by 9 CFR 113.53 in assays used to demonstrate that bovine, porcine, equine, and ovine raw materials are free of species-specific viruses.
Detection of Hemagglutinating Viruses—
A characteristic of certain viruses (referred to as hemagglutinating viruses) is that one or more of their viral proteins cause hemagglutination of one or more types of erythrocytes. Hemagglutination is an interaction between viral proteins or hemagglutinins and erythrocytes, leading to adhesion of the erythrocytes to surfaces, cells, and each other. This property forms the basis of two endpoint procedures that are employed in cell-based viral infectivity assays: hemadsorption and hemagglutination.
Hemadsorption is performed by adding a suspension of one or more erythrocyte types directly to the monolayer culture of detector cells. If viral proteins of a hemagglutinating virus are expressed from infected cell membranes, the susceptible erythrocytes will bind tightly to the cell membranes. Noninfected cells do not display this binding; therefore, the technique can be used to visualize a focus of infected cells against a background of uninfected cells. For this reason, this particular endpoint may display greater detection sensitivity than other assay endpoints, such as cytopathic effect or hemagglutination. In the advanced stages of infection, binding of erythrocytes to cells, to each other, and to open plastic surfaces in the culture vessel may be observed.
The hemagglutination procedure is performed on the conditioned medium and is essentially an evaluation for free virus or viral hemagglutinins in solution. An aliquot of the conditioned medium from a detector culture is combined, in a microwell plate having v-bottomed wells, with one or more types of erythrocytes. After an appropriate amount of time the plates are evaluated. Absence of hemagglutination is reflected by a well-defined pellet (button) of erythrocytes sedimenting to the bottom of the well. In comparison, hemagglutination is reflected by the absence of a button, or by a button with irregular shape. Scoring the latter represents an opportunity for operator subjectivity. In addition, this endpoint can be considered the least sensitive in detection assays, because it is dependent on achieving a sufficient concentration of viral hemagglutinins in solution. For these reasons, hemadsorption is typically viewed as the more useful and reliable of the techniques for detecting hemagglutinating viruses.
The responses obtained in detection assays using hemadsorption and hemagglutination endpoints are highly dependent on the virus being assayed, as well as on the types of erythrocytes used. Many of the viruses of concern to the biotechnology industry do not cause hemadsorption and hemagglutination, or their hemagglutinins react with red blood cell types not commonly used in detection assays.
Detection by Immunofluorescent Antibody (IFA) Staining—
Certain cell-based viral detection assays are intended to detect specific viral entities and are required for raw materials tests derived from bovine-, porcine-, equine-, and ovine-derived materials (9 CFR 113-53). In order to achieve this, the assays must be optimized with respect to host cell selection, study design, and sample preparation. Specificity of detection is also conferred through use of IFA staining techniques. Primary antisera or monoclonal antibodies directed against the viral antigens of interest are used, either in direct staining applications or in conjunction with a fluorochrome-conjugated secondary antiserum. The immunostained detector cell monolayers are then visualized with an epifluorescence microscope to reveal the presence of reactive infected cells.
Design of Cell-Based Viral Assays
Viral Detection Assays
Detector cell cultures are seeded and allowed to incubate for the appropriate amount of time. Viral samples may be inoculated directly into the medium of the mitotic phase of cell culture, but more typically the medium is removed from the overnight detector cell culture and is replaced with the test sample. For the latter method, the test sample must therefore be approximately isotonic, and cytotoxic agents such as selection agents must be maintained within levels tolerable to the detector cell. For test samples comprised of live cells, the sample must be subjected to freeze-thaw before inoculation in order to lyse the sample cells. If the cells are not lysed, a cocultivation involving the detector and sample cells will result. The latter is part of the design for cocultivation assays. But for most of the cell-based infectivity assays such a cocultivation is not intended and could adversely impact the sensitivity of the assay. The test sample is typically allowed to adsorb to the detector cell monolayers for an appropriate amount of time. It is then removed and replaced with the growth medium suitable for the detector cell. Once inoculated, the detection assay involves incubation of the detector cells for a prescribed amount of time, with periodic refeeding as necessary. Endpoint evaluations as described above are performed according to the study design. Variations of the given procedure may be used for exposing detector cells to raw materials. For raw materials, detector cell exposure may consist of incorporation of the test materials into the growth medium used to maintain the detector cells throughout the assay.
Viral Titration Assays
Viral titration assays are designed to generate quantitative information about the virus of interest. These assays do not quantify absolute numbers of viral particles; rather, the results are expressed in terms of infectious units. An infectious unit is the amount of virus required to establish a productive infection, and several categories of titration assays are used. They vary on the basis of the endpoint used to demonstrate infection and include viral plaque titration or plaque-forming units (units: PFU per mL); 50% tissue culture infectious dose (units: TCID50 per mL); and 50% fluorescent antibody infectious dose (units: FAID50 per mL). The amount of a hemagglutinating virus in a sample can also be expressed in terms of hemagglutinating titer (units: endpoint dilution, i.e., the greatest dilution of the sample which still results in a positive hemagglutination, or HA, response).
Regardless of the endpoint used, a typical titration assay design consists of a sequence of sample dilutions based on 0.5 log10 or 1 log10 increments. The various dilutions of the sample are then applied to an appropriate number of replicate permissive detector cell monolayers. After a suitable incubation time, the monolayers are scored directly for cytopathic effect (TCID50 assay), fixed and processed for immunostaining and scored for reactive cells (FAID50 assay), or overlaid with agarose and processed for plaque generation PFU assay. TCID50 and FAID50 titers are typically calculated using published formulas, such as Spearman-Kärber and Reed-Muench. Assay controls for such quantitative assessments should routinely include a reference sample of known potency.
Detection of Retroviruses
Retroviruses represent a special case for cell-based viral detection assay because of the occurrence of retroviral infection in the absence of responses to the typical endpoints discussed. In order to detect retroviruses, scientists can employ a number of different endpoints. A list of commonly employed indicator cell lines and associated endpoints, and their application in retrovirus detection assays, is provided in Table 4.
Table 4. Indicator (Detector) Cell Lines Used in Retrovirus Infectivity Testing
| Cell Line | Assay Type | Endpoint(s)a | Target Virus |
|---|---|---|---|
| SC-1/XC | Isolation/direct detection | Plaques (XC); RT | Ecotropic murine retrovirus |
| Balb/C | Isolation/direct detection | Plaques (XC); RT | B-Tropic murine retrovirus |
| NIH/3T3 | Isolation/direct detection | Plaques (XC); RT | N-Tropic murine retrovirus |
| Mink Lung | Isolation | RT; mink S+L- | Xenotropic, amphotropic murine retrovirus |
| Mink S+L |
Direct detection | Foci | Xenotropic, amphotropic murine retrovirus |
| Feline S+L |
Direct detection | Foci | Xenotropic, amphotropic murine retrovirus, gibbon ape leukemia virus (GALV), and RD-114 feline retrovirus |
| Mus dunni | Isolation, cocultivation | RT; S+L |
Murine retroviruses and retroviruses infectious to humans |
| QT6b | Isolation, cocultivation | RT | Avian retroviruses |
| RD | Isolation, cocultivation | RT; S+L |
Retroviruses infectious to humans |
| MRC-5 | Isolation, cocultivation | RT; S+L |
Retroviruses infectious to humans |
| WI-38 | Isolation, cocultivation | RT; S+L |
Retroviruses infectious to humans |
| 293 | Isolation, cocultivation | RT; S+L |
Retroviruses infectious to humans |
| A549 | Isolation, cocultivation | RT; S+L |
Retroviruses infectious to humans |
| Raji | Isolation, cocultivation | RT | Retroviruses infectious to humans |
| Human PBMC | Isolation, cocultivation | RT, HIV p24 EIA | HIV and other human retroviruses |
|
a
The endpoint assays include the following: RT, reverse transcriptase; PERT, product enhanced reverse transcriptase (including also product enhanced reverse transcriptase and real-time quantitative product enhanced reverse transcriptase assays); and EIA, enzyme immunoassay.
b
A quail cell; primary fibroblast cultures of chicken or turkey origin are also sometimes used.
|
|||
Retroviral infection is dependent on the presence of receptors on the host cell membranes. The presence of such receptors confers host cell tropism.
Design for Retrovirus Infectivity Assays
Two types of infectivity assays are used, depending on the nature of the test material. For materials other than intact cells, the test material is inoculated onto one or more of a variety of detector cells, and the latter are then passaged as required to amplify any virus present. Because of the nature of retroviral replication, cytopathic effects typically do not occur during infection, although there are some exceptions. Before the first subculture and at the end of the final passage, one or more endpoint assays are employed to detect the presence of a retrovirus. For test materials in the form of intact cells, detector cells are seeded and subsequently inoculated with the test cells, resulting in a cocultivation. The cultures are passaged five or more times. Before the first subculture and at the end of the final passage, one or more endpoint assays are employed to detect the presence of a retrovirus.
Endpoint Assays for Retrovirus Detection
Endpoint assays may be classified as direct, which lead to distinct morphological changes in the detector cells; or indirect, as measured by the detection of biochemical, molecular, or immunological markers for infection.
XC-Plaque Assay—
The XC-plaque assay was developed as a direct means of detecting infectious murine retroviruses. Detection of the retrovirus is accomplished by UV-irradiating the detector cells used to amplify the virus and overlaying the irradiated detector cells with a specific rat cell (XC). The presence of infectious murine retroviruses in the detector cells is reflected by the formation of distinctive syncytia in the XC monolayer, which are easily visualized when the cultures are fixed and stained with a suitable dye such as crystal violet.
The N/B tropism of an ecotropic murine virus may be determined by inoculating Balb/c and NIH Swiss detector cells with the isolate, performing one or two passages on each cell line, and comparing the XC-plaque titer post passage to that determined for the initial isolate.
Mink and Feline S+L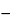 Focus Assays—
The S+L
Focus Assays—
The S+L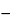 focus endpoint was developed to facilitate direct detection of infectious murine xenotropic and amphotropic viruses. The test sample may be inoculated directly into cultures of the S+L
focus endpoint was developed to facilitate direct detection of infectious murine xenotropic and amphotropic viruses. The test sample may be inoculated directly into cultures of the S+L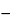 cells, or, alternatively, may be amplified first by inoculation into mink lung, human, or Mus dunni detector cells. Cell-free supernatants from the detector cell cultures are used to inoculate the S+L
cells, or, alternatively, may be amplified first by inoculation into mink lung, human, or Mus dunni detector cells. Cell-free supernatants from the detector cell cultures are used to inoculate the S+L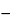 cells. The latter are infected with a sarcoma virus that is replication-defective, requiring the presence of a helper leukemia virus to render it capable of causing transformation of the host cell. The presence of infectious retrovirus virus in the detector cultures is reflected by the formation of characteristic focal areas of cell transformation in the S+L
cells. The latter are infected with a sarcoma virus that is replication-defective, requiring the presence of a helper leukemia virus to render it capable of causing transformation of the host cell. The presence of infectious retrovirus virus in the detector cultures is reflected by the formation of characteristic focal areas of cell transformation in the S+L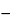 cells caused by the rescued sarcoma virus.
cells caused by the rescued sarcoma virus.
Detection of Retroviral Reverse Transcriptase—
Assays designed to measure reverse transcriptase (RT) activity are useful as an indirect detection method, because the enzyme is indicative of the presence of all retroviruses, whether infectious or not. The RT enzyme is encoded for in the retroviral genome and is used by the virus to transcribe genetic information in viral genomic RNA into proviral DNA.
Radiolabeled Nucleotide Incorporation Assay—
The earliest methods for measuring RT activity were based on the measurement of 32P- or 3H-labeled nucleotide incorporation into the complementary cDNA product, using an appropriate RNA template. Incorporation of radiolabeled nucleotide at levels higher than a predetermined threshold is interpreted as evidence of the presence of retroviral RT activity. The contributions of cellular DNA polymerases can be ruled out through use of a dual template assay (having both RNA and DNA templates) or inclusion of activated calf thymus DNA.
Product-Enhanced Reverse Transcriptase (PERT) or Quantitative PERT (Q-PERT)—
Polymerase chain reaction amplification has been used to increase the sensitivity of RT activity measurement. RT activity is detected by PCR amplification of complementary DNA, newly synthesized from an RNA template by reverse transcriptase. The assay may be performed with a gel endpoint (PERT) or as a quantitative assay (Q-PERT). This method has increasingly gained acceptance by regulatory agencies. For more details on PCR-based techniques, see the USP general information chapter Nucleic Acid Based Techniques—Amplification  1127
1127 .
.
Electron Microscopy—
Transmission electron microscopy (TEM) may be used to detect and enumerate viral particles within cells. In addition, the technique allows for differentiation of types A, B, C, and D retroviruses based on morphological considerations and can be used to localize viral particles within the cell. As a technique for identification of viruses (including RNA and DNA viruses in general), TEM of sectioned cells is extremely valuable. The cells, typically sampled during the log phase of growth, are pelleted by low-speed centrifugation, and the cell pellet is fixed with a suitable fixative. The fixed cell pellet is embedded, sectioned, stained, and observed with TEM. Size (diameter) of the particles, morphology, presence or absence of surface features such as envelopes and spikes, and location within the cell can be determined with this technique. Such information is important for the identification of a virus. However, failure to observe viral particles with this method does not conclusively demonstrate the lack of viral contamination in the sample.
Biological fluids may also be evaluated by TEM, primarily to determine particle size and concentration. The cell-free supernatant is subjected to ultracentrifugation to pellet any virus present. The resulting pellet is fixed with a suitable fixative. A predetermined number of grid spaces containing representative areas of thin sections of the pellet are evaluated for particles. The enumeration results obtained may show a high degree of variability, and failure to observe particles does not imply that none were present in the sample. Molecular (quantitative PCR and quantitative PERT) endpoints have also been used as alternative methods for estimation of viral particle load in samples.
Antigen-Capture Enzyme Immunoassay—
Specific viral proteins (e.g., HIV p24 antigen or avian leucosis viral envelope proteins) may be detected as a means of determining the presence of a retrovirus. Viral antigens are captured by specific antibodies coated onto microtiter plate wells and are detected by the addition of a second labeled antibody and appropriate substrate.
Assays Designed to Detect Specific Viruses
Additional methodologies have been developed to allow detection of specific viruses or groups of viruses. These types of assays are often used for raw material evaluation. In some cases, these specific assays were developed because the target viruses do not cause endpoint responses in the viral screening assays. In contrast to screening assays, specific virus assays are typically optimized for detection of the target virus or viruses. This optimization takes into account the lability of the virus, the host range, the possible endpoint responses elicited, and any special requirements of the target virus. The use of well-characterized viruses as positive controls in such assays provides assurance that the methodologies are suitable for the target virus or viruses. Spiking of the test sample matrix with the positive control virus enables the investigator to assess the potential for matrix interference and to assess the limit of detection for the method. Such considerations are not applicable to screening assays. Specific virus testing for bovine- and porcine-derived raw materials is discussed below. Evaluation of caprine, ovine, equine, canine, and feline raw materials is also stipulated in 9 CFR section 113.47. This section should be consulted with respect to the viruses of concern, and 9 CFR 113.52 should be consulted for methodology. A list of commonly employed indicator cell lines and their application in raw materials testing assays is provided in Table 2. A list of commonly employed indicator cell lines and their application in detection of specific viruses is also provided (see Table 3).
Detection of Bovine Virus Contamination—
Raw materials of bovine origin include such commonly employed medium components as fetal bovine and calf serum, serum albumin, collagen, thrombin, and trypsin. Each of these additives represents a route of entry for adventitious viral contaminants into a cell culture or manufacturing process. Requirements for evaluation of such materials ensure the absence of contaminating viruses.
For the details on testing for bovine serum and its derivatives, see future general chapter Bovine Serum  1024
1024 .
.
Detection of Porcine Viral Contaminants—
Raw materials of porcine origin include trypsin as well as other cell culture reagents. The specific porcine viruses of concern in the United States are stipulated in 9 CFR 113.47 and include porcine parvovirus, porcine adenovirus, transmissible gastroenteritis virus, and porcine hemagglutinating encephalitis virus. In addition, porcine raw materials must also be evaluated for the presence of bovine viral diarrhea virus (BVDV), reovirus, and rabies virus. Porcine tissues intended for xenotransplantation into humans also are routinely evaluated for the porcine endogenous retrovirus (PERV). The host cells typically used in the detection of porcine viruses are porcine testicle or porcine kidney, a bovine cell, and Vero cells. The methodology described in 9 CFR 113.52 is analogous to that for evaluation of bovine raw materials and includes provision for multiple subcultures, for Giemsa staining of fixed cells, for hemadsorption testing, and for use of specific immunostaining of fixed cells.
Cell-Based Detection of Murine Minute Virus—
Murine minute virus (MMV) is a mouse parvovirus that has been detected in biologics manufacturing involving Chinese hamster cell substrates. As with other parvoviruses, MMV represents a special case in that the virus is difficult to inactivate using typical cleaning agents and is capable of surviving for prolonged periods of time on surfaces. Cell-based assays for MMV involve detector cell lines that are especially susceptible to this virus, such as 324K (a human cell) and A9 (a murine cell). Optimization for detection of a parvovirus also includes provision for detector cell subcultures to remain in log-phase division for a significant portion of the incubation period. Endpoints for detection of MMV include one or more of the following: cytopathic effect, hemagglutination of mouse and guinea pig erythrocytes, immunostaining, and polymerase chain reaction.
Cell-Based Detection of Insect-Borne Viruses—
Insect-borne viruses include both viruses infectious only for insect cells (e.g., baculovirus) and those transmitted to mammalian cells via insect vectors (arboviruses). Detection of the former may be accomplished using an insect cell as a detector cell. Suitable substrates might include cells of Spodoptera, Trichoplusia, Drosophila, mosquito, or other insect origin. Such cells are typically cultured at lower temperatures (25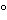 to 28
to 28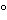 ) relative to mammalian cells, and many of these cultures are suspension or semiadherent at best. Endpoints may include cytopathic effect, electron microscopy, and PCR.
) relative to mammalian cells, and many of these cultures are suspension or semiadherent at best. Endpoints may include cytopathic effect, electron microscopy, and PCR.
Of more relevance to patient safety is the detection of arboviruses (insect-borne viruses infectious to animals and humans). This may be accomplished using a suitable mammalian detector cell. The Syrian hamster kidney cell (BHK-21) is a cell line that has shown susceptibility to a wide range of arboviruses. This cell line grows in a monolayer culture, and the endpoints that may be used include cytopathic effect, hemadsorption and hemagglutination, and PCR.
Cell-Based Detection of Human Cytomegalovirus—
Human cytomegalovirus (CMV) is a slow-growing virus of special concern for biologics produced using human cell substrates. It may be detected in cell-based assays using human diploid detector cells such as WI-38 or MRC-5, provided that sufficiently long durations of incubation are employed (28 or more days). The endpoints include cytopathic effects and immunostaining and/or PCR.
in vivo methods
Intact and susceptible animals may serve as potential host organisms for detecting viruses in test samples. In this case, viral proliferation in the tissues of the host animal may be reflected as adverse health effects (including death) that can be monitored and recorded. Viral detection assays based on intact animals are intended to complement in vitro assays, because some viruses that do not cause a response in the in vitro assays may be detectable in the animal systems (and vice versa). Viral safety studies employing live animals must be performed in accordance with applicable regional guidelines for the ethical use of animals, using laboratories that are accredited for the housing of the animals.
In Vivo Viral Screen—
The in vivo viral screen is used primarily for cell bank, viral seed stock, and viral vaccine testing and is considered to complement the in vitro virus screening assay. Multiple animal species, as well as multiple injection routes, are employed to provide a broad range of host tissues and possible responses. A list of commonly used host animals, routes of inoculation, and target viruses are shown in Table 5.
Table 5. In Vivo Viral Screening Assays
| Host Animal | Route of Inoculation | Target Virus |
|---|---|---|
| Suckling mouse | Intraperitoneal injection | Arboviruses |
| Intracranial injection | Coxsackie A and B | |
| Per os injection | Herpes simplex Type 1 and 2 | |
| Togaviruses | ||
| Junin | ||
| Herpes B | ||
| Adult mouse | Intraperitoneal injection | Rhabdoviruses |
| Intracranial injection | Togaviruses | |
| Per os injection | Lymphocytic choriomeningitis virus (LCMV) | |
| Guinea pig | Intraperitoneal injection | Rhabdoviruses |
| Intracranial injection | LCMV | |
| Lassa | ||
| Junin | ||
| Marburg | ||
| Ebola | ||
| Vaccinia viruses | ||
| Embryonated hens' eggs | Allantoic | Arboviruses |
| Yolk sac | Equine encephalomyelitis viruses | |
| Chorio-allantoic membrane | Herpes viruses | |
| Influenza | ||
| Mumps | ||
| Newcastle disease | ||
| Parainfluenza Types 1 and 2 | ||
| Rabies | ||
| Vaccinia | ||
| Variola | ||
| Lymphogranuloma venereum | ||
| Ornithosis |
Following injection of the test sample, each animal model is monitored for an appropriate period of time that allows for the observation of clinical signs of viral infection. Any abnormality is investigated to determine the cause of the effect.
The suckling mice are observed for an appropriate period of time. Pooled homogenates from any surviving animals are then passaged into additional litters of suckling mice. The latter are observed for an additional period of time.
The guinea pigs are observed for clinical signs of viral infection and for injection site lesions. Necropsy for gross tubercular lesions is performed for certain types of test samples.
Allantoic fluids from eggs can be tested for hemagglutination of chicken, guinea pig, and human type-O erythrocytes. Additional fluids are pooled for each treatment group (test article and control), and these are passaged (inoculated) into a new group of embryonated eggs. Following an appropriate incubation period (typically measured in days), the allantoic fluids are again tested for hemagglutination of chicken, guinea pig, and human type-O erythrocytes. Following injection by the yolk sac route, the eggs are incubated for at least 9 days and are assessed for viability. The yolk sacs are then harvested and pooled for each group (test article and control), and a 10% solution of the resulting material is inoculated by the same route into a new group of embryonated eggs. The eggs are again incubated for an appropriate period of time (days) and are assessed for viability.
in vivo assays intended to detect specific viruses
Some in vivo assays are designed to detect, if not specific viruses, at least specific sets of viruses. The antibody production assays use the production of a humoral immune response in susceptible host animals inoculated with test samples. Viral antibody-free animals of the various species are injected with the test sample. At the end of an appropriate incubation period, one or more of a variety of endpoint assays may be performed to detect the generation of a humoral antibody response in the animal sera. Production in the animal of antibodies directed against a specific virus provides evidence of the presence of viral antigen or infectious virus in the test sample. This type of assay is typically used to ensure that rodent cell banks and viral seed stocks are free of adventitious viruses. Three antibody production assays, along with the route of injection and target viruses, are summarized in Table 6.
Table 6. In Vivo Antibody Production Assays
| Antibody Production Assay | Route of Injection | Target Virus |
|---|---|---|
| Mouse antibody production (MAP) assay | Intranasal | Ectromelia |
| Intraperitoneal | Hantaan | |
| Intracranial | Mouse K | |
| Lactate dehydrogenase elevating virus | ||
| Lymphocytic choriomeningitis virus (LCMV)* | ||
| Murine minute virus | ||
| Mouse adenovirus | ||
| Mouse cytomegalovirus | ||
| Mouse encephalomyelitis virus type II | ||
| Mouse antibody production (MAP) assay | Mouse hepatitis virus | |
| Epizootic diarrhea of infant mice | ||
| Pneumonia virus of mice | ||
| Polyomavirus | ||
| Reovirus type 3 | ||
| Sendai | ||
| Mouse thymic virus | ||
| Hamster antibody production (HAP) assay | Intranasal | Lymphocytic choriomeningitis virus (LCMV)* |
| Intraperitoneal | Polyomavirus | |
| Intracranial | Reovirus Type 3 | |
| Sendai | ||
| Simian virus 5 | ||
| Rat antibody production (RAP) assay | Intranasal | Hantaan |
| Intraperitoneal | Kilham rat virus | |
| Intracranial | Mouse encephalomyelitis virus type II | |
| Polyomavirus | ||
| Reovirus type 3 | ||
| Sendai | ||
| Toolan's H1 virus | ||
| Rat coronavirus/sialodacryoadenitis virus | ||
|
*
A group of test sample–injected mice is challenged with a known lethal dose of authentic LCMV. If this group of mice does not die from the challenge dose, a second group with twice the number of mice is used for a repeat challenge. If there are survivors in this group, the test sample is considered positive for LCMV.
|
||
Considerations for Validation, Matrix Qualification, and Quality Control of Cell- and Animal-Based Test Systems
Viral detection assays used to ensure the viral safety of human and animal therapeutics are expected to have undergone validation. The approach to the validation depends on the nature of the assay and associated regulatory compliance level.
Any assay should be sufficiently developed that it can be performed with an appropriate set of predetermined system suitability and acceptance criteria. These criteria usually include the use of relevant negative and positive controls but may also include requirements for linearity and meeting of a predetermined detection limit. The results constituting a positive or negative response in the assay should be established prior to execution of the validation. An assay used under GMP compliance is expected to have been validated according to appropriate guidelines.
An assay used to ensure the safety of a commercially marketed biological product must be further characterized for suitability in the presence of the specific product matrix. The matrix qualification study should address the potential for specific interference with the viral detection endpoints used in the assay, and typically involves spiking of one or more model viruses into the product matrix at levels approaching the limit of detection to ensure the absence of interference.
For quantitative detection assays, the detection limit should be probed. This usually involves spiking of the model virus(es) at decreasing amounts into medium or the product matrix. The lowest spiking level of the virus reliably detected is used as an approximation of the actual limit of detection of the assay. Experimental error for these cell culture-based assays is usually expected to be in the range of 0.5 to 1 log10. The determination of a detection limit is less meaningful for limit tests and viral screening assays in general. For the latter, knowledge of the detection limit for one virus does not imply a similar limit for another virus. Since screening assays are not optimized for a specific virus, the limit of detection for the assay can vary greatly from one virus to another.
Animal-based viral detection systems are generally not subject to the requirements for validation, matrix qualification, use of positive controls, and determination of detection limit that regulatory agencies expect of cell-based and biochemical tests. The use of animals for safety testing is subject to the regional guidelines for the ethical use of animals, and the kinds of activities listed above generally are not considered appropriate use of animals. However, negative control animals are included in these assays, and retrospective validation or gap analysis based on historic incidence of system suitability failures or positive findings is sometimes possible.
DETECTION OF VIRAL COMPONENTS
Direct detection of viral components can provide a direct measurement of viral levels in a sample preparation. It has also become primarily important for detection or identification of viruses in biological products or in the raw materials used in their manufacture. Systems capable of identifying components unique to specific phases associated with viral latency and replication are now available. During interaction with their host cells, viruses may incorporate modified host molecules during the production of new intact virus particles, or they may induce discernable changes in host cell makeup or function.
Most immunological methods and reagents currently available detect the constituents of intact virions. It is the relative abundance of these proteins that makes them most amenable to the development of antibody-based reagents. Abundance also makes them optimal targets for detection of the virus. Recently developed targeting and detection reagents are aimed at minor viral components that may be found only during specific phases of replication. These allow a more detailed analysis of the stage of viral infection. The basic methodology for the detection of viral antigens is well established, but more recent innovations in materials and reagent development have broadened its application.
Developments in the targeting and detection of viral nucleic acid components have led to enzyme-based systems for the amplification of nucleic acids in vitro and in situ (see Nucleic Acid-Based Techniques—Amplification  1127
1127 ). The potential specificity of this detection method allows the examination of biological systems with a high degree of confidence for the presence or absence of a specific targeted virus. A wide variety of reagents, technology platforms, and methodologies are available. The aim of this section is to elaborate on the most common practices and platforms used in the detection of viral components.
). The potential specificity of this detection method allows the examination of biological systems with a high degree of confidence for the presence or absence of a specific targeted virus. A wide variety of reagents, technology platforms, and methodologies are available. The aim of this section is to elaborate on the most common practices and platforms used in the detection of viral components.
Sample Selection and Preparation for the Detection of Viral Components
This subsection addresses general considerations for various types of test samples and the most common assay targets (viral proteins and nucleic acids). The target proteins may have varying levels of posttranslational modification (e.g., glycosylation, phosphorylation), and the target nucleic acids may be either RNA or DNA, single or double stranded. Therefore, it is important to have a basic understanding of the physicochemical nature of the virus under study so that the sample handling procedures support the detection of the target component. For detailed considerations regarding the extraction of nucleic acids, see the USP general information chapter Nucleic Acid-Based Techniques—Extraction, Detection, and Sequencing  1126
1126 .
.
Types of Samples
Cellular—
When viral components are associated with intact cells, samples must be generated either as whole cell lysates or as subcellular fractions. Maintaining the temperature of the test samples at or near freezing (0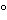 to 4
to 4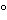 ) during processing and limiting the time that samples are held in an unfrozen state will reduce the potential for loss of target antigens and nucleic acids due to cytosolic enzymes (proteases, nucleases) present in the cell lysates. Reagents that can inactivate or limit the activity of such enzymes may be used to prevent degradation of the target components, especially when exceptionally labile samples must be handled at room temperature. Centrifugation of intact cells allows for some additional manipulation of the sample matrix. The growth medium can be discarded and the cells suspended in a buffer formulated to enhance the recovery and detection of the targeted viral component. Collection and storage parameters should also account for the presence of cellular DNA. This can increase the viscosity of the sample, rendering it difficult to pipet.
) during processing and limiting the time that samples are held in an unfrozen state will reduce the potential for loss of target antigens and nucleic acids due to cytosolic enzymes (proteases, nucleases) present in the cell lysates. Reagents that can inactivate or limit the activity of such enzymes may be used to prevent degradation of the target components, especially when exceptionally labile samples must be handled at room temperature. Centrifugation of intact cells allows for some additional manipulation of the sample matrix. The growth medium can be discarded and the cells suspended in a buffer formulated to enhance the recovery and detection of the targeted viral component. Collection and storage parameters should also account for the presence of cellular DNA. This can increase the viscosity of the sample, rendering it difficult to pipet.
Tissue Culture Supernatant—
Depending on the stage of the infection and the type of virus involved, the conditioned medium may represent a preferred test sample. An advantage is that the presence of cellular debris can usually be reduced through use of a low-speed centrifugation (clarification) step. The main disadvantage is the potentially low concentration of the analyte, and therefore concentration of the sample may be required.
Process Intermediate (Unprocessed Bulk Harvest)—
In general, viral safety lot release testing is done at the bulk harvest stage prior to any purification. This is true regardless of whether the assay detects infectious virus or viral components. The presence of host cell DNA may need to be assessed in the case of biologics manufactured in animal cells.
Sample Stability and Matrix Effect
Sample stability is a key element in the successful detection of viral components. Protein structure can be altered by numerous environmental factors, including pH, ionic strength, solvents, detergents, temperature, and free radicals. In addition, complex biological matrices frequently contain proteolytic enzymes that can alter or destroy key antigenic features of a protein or peptide. Sample collection, storage, and handling must allow maintenance of the antigenic features targeted by reagent antibodies.
Conformational changes affecting the opportunity for antigen detection are difficult to address. Depending on the reagents required for detection, conformational changes may be required for antigen detection. For example, if antibodies are produced for an antigen detection system using native viral antigen, then unmasking and maintaining the conformation of the antigen throughout the sample preparation is essential. Conversely, if peptide fragments are used to produce antibody, then a denaturation step may be required to allow for effective antigen detection.
Conditions associated with sample preparation must be investigated in a combinatorial fashion whereby one parameter or component is varied while all others remain fixed. In this way, the formulation of lysis and processing buffers can be optimized for pH, ionic strength, and types of detergents and denaturants. The sample preparation steps must condition the targeted antigen in order to obtain the form most readily recognized by the reagent antibody.
The stability of nucleic acids in test samples is largely affected by nuclease activities present in the sample and the degree of protection provided by the intact structure of the virus particle. Encapsidated nucleic acids are particularly stable as long as the integrity of the capsid is maintained. Viral capsids are vulnerable to proteolytic digestion. Virus particles stored at ambient temperature as part of a complex biological matrix are especially susceptible to degradation by proteases. Storage at refrigerated temperatures (2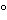 to 8
to 8 ) for short periods of time or at temperatures below freezing can be used to limit proteolytic activity. When samples are stored at frozen temperatures, freeze-thaw cycles should be limited. Under conditions where an individual sample must be accessed multiple times, preparation of aliquots is advisable.
) for short periods of time or at temperatures below freezing can be used to limit proteolytic activity. When samples are stored at frozen temperatures, freeze-thaw cycles should be limited. Under conditions where an individual sample must be accessed multiple times, preparation of aliquots is advisable.
Sample Collection—
In a biotechnology setting, sample collection is dictated by sampling plans that are established to meet regulatory requirements. Nucleic acid testing in association with an amplification step has the potential of detecting a virus at the earlier stages of infection. The use of nucleic acid amplification methods reduces the dependence on timing and the amount of material required, because the amplification process effectively boosts assay sensitivity by increasing the amount of target relative to background.
General aspects of nucleic acid sample preparation and stability are discussed in Nucleic Acid-Based Techniques—Extraction, Detection, and Sequencing  1126
1126 . The following section specifically addresses the unique aspects of viral sample selection and preparation.
. The following section specifically addresses the unique aspects of viral sample selection and preparation.
Sample Storage—
Conditions for sample storage should be consistent with maintaining the antigenic properties of targeted viral proteins and/or preserving the nucleic acid content of the sample. The duration and temperature of storage is dictated also by cycle times associated with testing.
Immune Complex Disruption—
The masking of antigen epitopes may occur when other proteins associate at or near the epitope targeted by reagent antibodies. For viral antigens this may occur when the antigen comprises the structural component of the virus. Such viral antigens are likely to retain strong affinities for other viral proteins or the ability to exist in multimeric form under normal conditions for detection. Another obstacle for epitope recognition is the naturally occurring immune complex when reagent antibodies have been developed to detect native protein in a blood plasma matrix. Immune complexes consisting of viral antigens and host antibodies are normal in such physiologic samples. The stronger the host immune response, the more likely masking of antigen due to immune complex formation will occur. Methods aimed at the preparation of blood and plasma samples for detection should address the presence of preexisting immune complexes and incorporate steps designed to disrupt such complexes to improve the opportunity for viral antigen detection.
Detection of Viral Antigens
Viral capsid proteins are common targets for antigenic detection methods. Structural proteins that make up the framework of the viral core are often some of the most abundant viral proteins produced during viral replication. In nonenveloped viruses, the core structural proteins are likely to provide the dominant antigenic features. When the virus is enveloped, proteins associated with the envelope often provide key antigenic features. This section examines methods commonly used to detect viral antigens and addresses considerations aimed at optimizing formation of the appropriate immune complex.
Assays used for the detection of viral antigens
Immunologic methodologies used to detect viral antigens are based on the specificity and affinity of the antibody and viral antigen interaction. Of the various platforms available, one commonly employed in viral safety testing is immunofluorescent antibody staining. This lends some degree of viral specificity to the cell-based methods described in the first part of this chapter. Other techniques include enzyme-linked immunosorbent assay (ELISA), radioimmunoassay, and Western blotting. The principles and general methods for these assays will be described in the USP general information chapters Immunological Test Methods—General Considerations  1102
1102 , Immunological Test Methods—Reagent Development
, Immunological Test Methods—Reagent Development  1103
1103 , Immunological Test Methods—Immunoassay Methodologies
, Immunological Test Methods—Immunoassay Methodologies  1104
1104 , and Immunological Test Methods—Assay Design, Quality Control, and Data Analysis
, and Immunological Test Methods—Assay Design, Quality Control, and Data Analysis  1105
1105 , being prepared for future publication. The following sections will generally address aspects specific to their use in virology.
, being prepared for future publication. The following sections will generally address aspects specific to their use in virology.
Immunofluorescence Assay—
The immunofluorescence assay (also referred to as immunofluorescent antibody staining) is used to detect viral proteins expressed in various cellular compartments. Since the technique can detect viral antigens within single cells, it confers a high degree of sensitivity and enables infection to be detected at a very early stage. The technique is often employed as a detection endpoint for cell-based viral infectivity assays to provide additional sensitivity and specificity (see tests for raw materials as described in 9 CFR 113.53). In addition, the technique is employed for verifying the identity of viral stocks and is useful as a means of identifying viruses detected in viral screening assays.
Enzyme-Linked Immunosorbent Assay (ELISA)—
ELISA is best suited for detection of soluble antibodies and antigens in a variety of test samples. Sensitivity, quantification, robustness, ease of experimentation, and readily available inexpensive reagents make it adaptable to a high throughput environment. For viral antigen detection, a sandwich ELISA assay is commonly used. A viral antigen specific antibody (preferably a monoclonal antibody with high affinity) is first immobilized onto a solid phase. The test sample is then incubated for a predetermined period under appropriate conditions. After washing, a second antibody that recognizes the viral antigen is incubated. The second antibody is linked to either an enzyme or a chromomeric reagent that emits signal with an appropriate substrate and can be recorded with an appropriate instrument. The assay can be performed in a qualitative or a quantitative manner. For a qualitative ELISA assay, sufficient replicates of both positive and negative control samples are required in order to determine the appropriate cut-off value and the assay acceptance criteria. A mean value of a test sample that is equal to or greater than the cut-off value is considered positive. For a quantitative ELISA assay, an additional standard curve with a positive reference standard material of known quantity must be established. The number of replicates should be adequate to determine the assay variation and linearity. The quantity of test sample can be calculated against the standard curve.
Radioimmunoassay (RIA)—
The radioimmunoassay is a versatile quantitative immunoassay that can be used to detect substances including viral antigens and antibodies. It even can be applied to nonprotein molecules as long as an antibody that specifically binds the test substance is available. Radioimmunoassays can be customized in different formats to suit specific test requirements. Many of the considerations taken into account with other immunological assays are applicable to the RIA. In general, radioimmunoassays can be divided into two major categories: solution (homologous) and solid-phase radioimmunoassays. Both methods have been successfully used to detect and quantify a variety of viral antigens or components of viruses, such as hepatitis A, B, and C; human and murine retroviruses; adenovirus; avian C-type virus; rubella virus; and respiratory syncytial virus.
Western Blotting (Immunobloting)—
Western blotting, also known as immunoblotting, is used to identify specific antigens in the presence of other, potentially cross-reactive antigens. In this case, the specificity required is obtained by combining the antigen-antibody reaction with some form of separation (typically electrophoresis). Depending on the visualization methods employed (including digital methods such as densitometry), this method can be quite sensitive and even semiquantitative. One advantage of this approach is that in addition to detecting the antigen using an immunological approach, data on the approximate mass of the target protein may be obtained. A potential caveat is that most test proteins are denatured during this procedure and that antibody that depends on epitope conformation may not recognize the linear epitopes.
detection of viral nucleic acids
The detection of viral nucleic acids provides another route for the determination of viral loads and for establishing the identity of a contaminant. Nucleic acids, like protein antigens, are essential components of viruses, and detectable quantities are usually indicative of viral presence. Detection assays can be designed and developed in some cases to parse viremia into phases, especially when the differentiation of nucleic acids along functional forms and configurations can provide clear insight into viral activity. Assays can be designed to determine whether viral DNA has been integrated into the host genome or still is encapsidated. Early viremia may be detected as viral mRNA transcripts prior to the accumulation of detectable viral particles. Nucleic acids may be the only detectable viral component of viruses that do not replicate well in tissue culture systems. Such systems may fail to produce mature virus particles, but the detection of viral transcripts can provide insight into whether the virus has the ability to infect the cell. Nucleic acid testing represents the most useful endpoint for the detection of certain viruses failing to cause responses using typical endpoints.
sample preparation: special considerations for nucleic acid testing
The degradation of nucleic acids in samples can be limited through proper handling and storage practices and even enhanced by closely linking sample collection and preparation steps. In addition to preparing nucleic acids for further processing, denaturation is an important step toward stabilizing nucleic acids where storage temperatures extend above 0 .
.
Denaturation and Dissociation of Virions (Viral Lysis)—
Chaotropic detergents and salts can be important agents for disrupting and removing viral proteins that make up the viral capsid. Their addition can provide a useful first step when concentration of virions is not necessary or even possible. In sufficient quantity they rapidly denature the entire contents of a biological sample, essentially fixing nucleic acid content through the inactivation of nucleases and other proteins that may affect sample stability. Saturated solutions containing guanidium salts, such as guanidine hydrochloride or guanidine isothiocyanate, are commonly used for the dissociation of viral nucleic acids from protein components. These solutions may be used alone or in combination with ionic detergents and other denaturants such as phenol. The main advantage of guanidinium salts is that they are readily removed during the concentration of viral nucleic acids using ethanol or isopropanol. Urea may also be used as a mild denaturing agent, although it does not perform as effectively as guanidine in its ability to disrupt virus particles.
Deproteinization—
The removal of proteins during the processing of samples for the detection of viral nucleic acids is helpful in ensuring the reproducibility and robustness of assays, particularly those that rely on amplification to detect exceptionally low quantities of nucleic acids. Several strategies may be used to facilitate deproteinization of the sample; they are discussed in Nucleic Acid-Based Techniques—Extraction, Detection, and Sequencing  1126
1126 .
.
Recovery of Viral Nucleic Acids—
Separation and recovery of extracted viral nucleic acid are important steps in the testing of nucleic acids. Nucleic acid yield and purity obtained at this step are critical determinants of assay robustness. Poor nucleic acid recovery and limited purification may inhibit amplification and detection reaction resulting in poor assay sensitivity. The development of high-yield, high-purity recovery steps is an important goal in the optimization of nucleic acid detection methods. Details on the general aspects of extraction and detection of nucleic acids are discussed in Nucleic Acid-Based Techniques—Extraction, Detection, and Sequencing  1126
1126 . However, if no inhibitory components are identified in the system, direct testing on an aliquot of a sample may increase the assay sensitivity.
. However, if no inhibitory components are identified in the system, direct testing on an aliquot of a sample may increase the assay sensitivity.
Isolation of Viral DNA—
Integrated viral genomes must be recovered and processed along with the genome of the host cell. The viral genome is analyzed within the context of the host genome, and therefore similarities between virus and host genomes must be accounted for to ensure that assay results are specific for the virus and do not simply reflect the presence of cross-reacting host genome sequences. Viral genomes that exist as episomal entities require similar consideration, but there may be opportunities during sample preparation to limit the amount of host nucleic acid present in the preparation. Sedimentation gradients or silica-based separation chromatography can sometimes be used to enrich episomal nucleic acids through size-related exclusion or partitioning of processed nucleic acids.
The recovery of encapsidated viral genomes may allow for larger amounts of material, especially if the target virus has accumulated in large quantities in the system being monitored. When the amount of material available for recovery is low or the processing of large amounts of the material is impractical, the process of isolating and conditioning the virus particles, and subsequently the encapsidated genome, must be compatible with the assay system that will be used. Portions of the viral genome may need to be amplified, or the genome itself may be captured in an elaborate process that allows detection through the generation of an amplified signal. A standard method for isolating viral genomes may need to be modified appropriately to ensure that materials used in the preparation do not interfere with steps conducted later in the process.
Viral genomes that consist of RNA are typically converted to a DNA intermediate that is easier to handle and store. Methods aimed at the isolation of viral genomes consisting of RNA, however, require similar considerations concerning the localization of the genome: replicating RNA viral genomes may be recovered as part of a preparation of host total RNA or even mRNA if the genome contains polyA sequences. RNA lends itself more effectively to hybrid capture methods during isolation, and hybrid capture can be used to enrich RNA preparations specifically for RNA viral genomes. RNA genomes can be extracted from virus particles in much the way that DNA viral genomes are extracted. RNA genomes are converted to complementary (cDNA) sequences using retroviral RT. Storage is of greater concern for naked viral RNAs. Storage of RNA usually requires temperatures below  20
20 .
.
detection of viral genome versus viral transcripts
Viral genomes exist as either DNA or RNA, or sometimes both: in the case of retroviruses the integrated genome is DNA, whereas the encapsidated form is RNA. The ability to differentiate among the various forms of viral nucleic acids can help to elucidate the course of specific viral infections. Assays for nucleic acid activity can differentiate readily between integrated and encapsidated genomes when the form of the viral nucleic acid varies between states, as in the case of retroviruses. Incorporation of specific nucleases into the assay methodology can be used to reduce or eliminate one form over the other. If viral genomes are known to integrate at specific sites within the host genome, primers and probes can be developed around the integration site and incorporate significant elements of both host and viral genomes. Some viral mRNAs contain splice sites, and the differentiation of spliced nucleic acid sequences from unspliced sequences creates a unique mechanism for determining the status of nucleic acid localization and infection.
Characterization of DNA Viral Genomes—
Methods for the recovery and preparation of viral genomes for characterization depend on the state of the viral genome. If the genome has been incorporated into a cellular compartment, the recovery and preparation strategy must take into account the cellular components that make up the sample matrix. If the viral genome targeted for analysis is the encapsidated form, the methods must focus on recovery of the virus particle and must include additional steps aimed at extracting the nucleic acid from the individual particles. Identification and characterization of viral genomes require specific complementary nucleic acid probes and primers whose sequence will be dictated by available information about the sequence of the targeted viral nucleic acids and the type of assay that will be used. Determination of the sequence of the viral nucleic acid of interest usually provides the most unambiguous means for characterization. However, a number of methods can be used as simple indicators for the presence or absence of specific sequence-based characteristics. For example, melting curve profiles using short oligonucleotide sequences can be used to establish whether a specific viral genotype is present.
Identification and Genotype Analysis—
Nucleic acid testing is often used to identify viral isolates obtained from viral screening assays or to provide identity for viral stocks. Methods used for the identification of viral genomes are not unique to other applications in the field of molecular biology. Typically, an amplification step is required in order to achieve quantities for analysis. Amplified sequencing of the amplicons or application of a standard hybridization technique may be employed for more detail as to the nature of the amplified signal. For more details, refer to Nucleic Acid-Based Techniques—Amplification  1127
1127 .
.
Hybridization Techniques—
A variety of hybridization techniques are used to detect viral nucleic acid sequences, including Southern blot, Northern blot, DNAse/RNase protection, in situ hybridization, microarray technology, and other techniques. The description of these methods, which is well beyond the scope of this chapter, can be found in Nucleic Acid-Based Techniques—Extraction, Detection, and Sequencing  1126
1126 .
.
DETECTION OF ANTIBODIES TO VIRAL ANTIGENS
A variety of methods are available for detection and quantification of antibodies to viral agents, including neutralization, complement fixation, and immunoassays based on enzyme- or fluorescently labeled reagents. Although many details of the immunological methods mentioned above are beyond the scope of the chapter, this section addresses specific application aspects of viral antibody detection, including preparation and storage of test samples, common assay methods and platforms available, and specific examples of how these assays may be used to measure antibodies to specific viral agents.
Viral Structure Relative to Antigenic Composition and Selection of Antibody Assay
Mammalian viruses vary considerably in their nucleic acid content and thus the number of antigenic, virus-specific proteins produced.
Many viral proteins or glycoproteins are highly antigenic and induce a potent humoral immune response during natural infection, whether in humans or in animal models. In most cases, the immune system responds when the virus and its antigens appear in the extracellular fluid or on the infected cell membranes. The degree to which viral antigens are expressed is governed by the intracellular replication and protein synthesis of viruses in host organ tissues and by the several possible types of virus–host cell interaction. Antibodies produced as a result of natural viral infection are likely to represent the broadest response to antigens in their native state.
When selecting, developing, or evaluating an assay method for measurement of antibodies to viral proteins, the analyst must take into account the source of the antibodies and the method by which they were obtained or prepared. Details will be presented in Immunological Test Methods—Reagent Development  1103
1103 and Immunological Test Methods—Immunoassay Methodologies
and Immunological Test Methods—Immunoassay Methodologies  1104
1104 .
.
General Considerations Regarding Sample Preparation for Antibody Detection of Viral Antigens
Antibodies are relatively stable, but care must be taken to ensure the integrity of the test antibodies during sample preparation and storage. Serologic tests may be developed to measure antibodies to viral agents in unfractionated biological fluids. The possibility for matrix interference with the antibody detection method should be considered.
In general, biological test samples should be clarified by centrifugation or filtration, depending on their intended use. Serum samples should not be hemolyzed, lipemic, or icteric. In some cases the specimen should also be heat-treated to inactivate endogenous complement and other components.
Test samples should be processed as soon as possible. When it is necessary to store samples, most test samples should be stored at  20
20 for short-term storage and below
for short-term storage and below  80
80 for long-term storage. For all samples, the stability of the material needs to be assessed experimentally. Aliquots of appropriate volume should be prepared in accordance with test procedures to avoid unnecessary freeze-thaw cycles.
for long-term storage. For all samples, the stability of the material needs to be assessed experimentally. Aliquots of appropriate volume should be prepared in accordance with test procedures to avoid unnecessary freeze-thaw cycles.
Antibody Methods
This section discusses the primary methods for detecting antibodies directed against viral antigens. The assay methods often include a variety of alternative formats for the detection of antibody. Only the more commonly used formats for antibody detection are discussed in this section. Some methods, including fluorescent antibody assays and enzyme immunoassays, are widely applicable to the detection of antibodies to many different viral agents; others are limited to selected viruses having certain properties (e.g., hemagglutinins).
Immunofluorescence Microscopy for Antibody Detection
When fluorescein isothiocyanate (FITC) is chemically coupled to an antibody molecule, the resulting FITC-labeled antibody can be used as a secondary antibody probe to detect the presence of a primary, virus-specific antibody bound to a virus-infected cell on a microscope slide (indirect immunofluorescence).
The indirect immunofluorescence or indirect fluorescent antibody (IFA) assay is one of the most basic and useful methods for detection of antibodies to viruses. The assay can be used to detect both virus-specific IgG- and IgM-class antibodies. When the assay is used to detect IgM antibodies, it usually requires the physical removal or inactivation/binding of IgG-class antibodies. In the absence of this step, the presence of IgM-specific antibody may be masked by excess IgG-specific antibody competing for primary binding sites on the substrate surface. IFA assays may be qualitative or quantitative.
The IFA for antibody to viral agents requires the use of virus-infected cells expressing viral antigens in cellular membranes. Viral stocks are prepared, titered, and used to infect permissive cells in tissue culture. The cells are harvested at appropriate times, washed, and spotted onto multiwell microscope slides at an appropriate density. Control slides are also prepared with noninfected cells. The slides are allowed to air-dry and then fixed in cold acetone. The fixed slides can then be stored under appropriate conditions for extended time periods. The stability of the viral antigens over time should be confirmed.
The test article to be examined for the presence of virus-specific antibodies can be applied to the slide, followed by an appropriate secondary antibody conjugated with a fluorescent tag that can be visualized under a fluorescent microscope. IgG- or IgM-specific antibodies can be distinguished by using the appropriately prepared secondary antibody.
Reading and correctly interpreting endpoints of IFA slides for antibody detection requires an experienced analyst, particularly when cellular location and fluorescent-staining patterns are critical for a specific virus. Such interpretation requires the use of appropriate controls and scoring or intensity of fluorescence. This is highly dependent on the quality of reagents, the consistency of the fluorescent microscopy and light source being used, and the experience of the analyst.
Enzyme Immunoassay for Antibody Detection
The EIA and variations of it are the most widely used methods for the detection of viral antibodies in serum and other biological products. The most commonly used EIA for antibody detection is referred to as a noncompetitive solid phase EIA for antibody detection. The typical configuration of an EIA for antibody involves coating tubes or microwell plates with viral antigen(s), the addition of test serum or product to the tubes or wells, the binding of specific antibody in serum or product to antigen, and the detection of bound antibody by addition of a second antibody with binding affinity to the primary antibody, which is labeled to allow for its detection.
The assays can be specific to IgG- or IgM- class antibody or may detect total antibody. Assays for IgM may achieve improved specificity when performed as IgM-capture assays. These assays involve the use of plates or wells coated with anti-IgM antibody to capture total IgM in serum or product as a first step. Subsequently, viral antigen is added; it binds to the plate only if virus-specific IgM antibody has initially been captured, and it is detected by addition of a second labeled antibody specific to viral antigen. The assay is most often performed as a qualitative measure of the presence of an antibody for a specific virus. Sufficient replicates of both positive and negative control samples are required in order to determine the appropriate cut-off value and the assay acceptance criteria. A mean value of a test sample equal to or greater than the cut-off value is considered positive.
Complement Fixation Test
Complement fixation has selective value in allowing for simultaneous assay of antibodies to a wide variety of viral agents. The procedure involves multiple variables consisting of two pairs of antigen–antibody reactions. The first reaction, between a known virus antigen and a specific antibody in the test sample, takes place in the presence of a predetermined amount of exogenous complement. The complement is removed by the antigen–antibody complex. The second antigen–antibody reaction consists of sheep red blood cells (SRBCs) and hemolysin (antibody against SRBC). When this indicator system is added to the reaction mixture, the sensitized SRBCs will lyse only in the presence of free complement. The extent of lysis of SRBCs is inversely correlated with the amount of the antibody in the test article.
The experimental procedure involves the optimal titration of concentrations of hemolytic serum, complement, and viral antigen, using chessboard format. If used as the test sample, human serum should be inactivated at 56 for 30 minutes to inactivate the endogenous complement activity. A number of important controls must be run along with the test, and results must be within limits before the test can be properly interpreted. These include the sensitivity of SRBCs to lysis and complement concentration used. The relative amount of virus-specific antibody present can be determined by testing serial dilutions of the serum or product. The complement-fixing titer is the reciprocal of the highest dilution that prevents 50% hemolysis.
for 30 minutes to inactivate the endogenous complement activity. A number of important controls must be run along with the test, and results must be within limits before the test can be properly interpreted. These include the sensitivity of SRBCs to lysis and complement concentration used. The relative amount of virus-specific antibody present can be determined by testing serial dilutions of the serum or product. The complement-fixing titer is the reciprocal of the highest dilution that prevents 50% hemolysis.
Neutralization for Antibody Detection
Neutralization for the measurement of antibodies to viral agents is still one of the most valuable assays available because of its high specificity and its ability to detect neutralizing antibodies. Neutralization is defined as the loss of viral infectivity through the binding of specific antibodies to viral coat proteins (or envelope glycoproteins) on the surface of the infectious viral particle. The assay may be used to measure the presence of antibodies to a known virus in a serum or product sample, or conversely to identify an unknown virus by using a serum or product sample containing known antibodies.
Before performing a neutralization assay to measure the presence of antibodies in serum or product, a known virus must first be grown and titrated in the test system in which the neutralization assay will be performed. For viruses prepared in cell culture, this usually involves inoculating susceptible cultures with relatively low multiplicity of infection (MOI; <1 PFU per cell) and harvesting the infected cells when about 50% to 75% cytopathic effect (CPE) is demonstrated. The virus preparation is then titrated by preparing serial multifold dilutions and inoculating replicate tubes or plate cultures with a fixed volume of the virus preparation. The endpoint of the titration is the dilution of the virus that will infect 50% of the cell cultures inoculated. This endpoint is said to contain one 50% tissue culture infective dose (TCID50) in the volume used. If the test system involves animal lethality, the endpoint is referred as one 50% lethal dose (LD50). The amount of virus used in the neutralization assay to follow is typically standardized to contain 100 TCID50 or LD50.
The test or host system used in neutralization assays is chosen on the basis of the specific virus to be tested and its ability to replicate in the system. The commonly used host systems include cell culture, embryonated chicken eggs, and mice. Cell culture is usually the preferred test system, because the viruses used in the neutralization assay usually readily replicate and produce CPE. Susceptible host cells are grown in monolayers in dishes or multiwell plate cultures. After the virus/neutralizing serum mixture is added, the cultures are overlaid with agar-containing medium to restrict spread of CPE and allow development of viral plaques. The prevention of plaque development is indicative of the presence of neutralizing antibody. Alternatively, neutralization can be performed in tube monolayer cultures or even in suspension tissue culture. Embryonated eggs may be used when the virus to be used or tested does not produce plaques in tissue culture systems. The route of inoculation and the endpoint depend on the virus.
Neutralization assays may be set up in various ways, depending on the specific virus of interest and the serum or product to be tested for neutralizing activity. In general, a fixed amount of infectious virus is preincubated with undiluted and serial dilutions of serum or product to be tested for neutralizing activity and separately with preimmune serum or control product; this approach is referred to as the constant virus–varying serum method. Following preincubation, the mixtures are separately injected or added to the test system. Reduction in infectivity between test and control serum or product is scored in various ways, depending on the test system. The endpoint of the assay is generally defined as the highest dilution of the serum or product that neutralizes one-half of the initial viral inoculum, as calculated by Reed-Muench or the Spearman-Kärber method.
The titer of neutralizing antibody in the test serum or product is the reciprocal of the highest dilution that completely inhibits CPE or other virus effect in the test system. This dilution is said to contain 1 neutralizing antibody unit per unit volume used in the titration. When a serum or product known to contain neutralizing antibody is used in an assay to determine the identity of an unknown virus, 20 neutralizing antibody units in a fixed volume are generally used in the assay. Positive and negative control sera must give expected reactivity in the assay.
Hemagglutination Inhibition (HAI)
A number of enveloped viruses, including the influenza and parainfluenza viruses, acquire protein receptors capable of binding RBCs (hemagglutinins) of various animal species on their surface as they bud through infected cell plasma membranes during viral maturation. In addition, some nonenveloped viruses such as adenoviruses and certain enteroviruses have hemagglutinin proteins in their outer capsid. This property allows for detection of a specific virus in a sample if a known specific antibody to the virus is available. Alternatively, the presence of antibody specific to the virus can be detected and quantitated by its ability to inhibit hemagglutination. This is the principle of the hemagglutination inhibition (HAI) test.
The HAI test for antibody is performed by making serial dilutions of the specimen to be tested and mixing the dilutions with a fixed amount of the virus or specific viral hemagglutinin protein in a tube or microtitration plate format. Indicator RBCs from the appropriate animal species are added, the suspension is mixed, and the tubes or plates are allowed to stand for a predetermined period. If specific antibody is present, the virus will bind and the RBCs will not agglutinate; they will settle to the bottom of the tube or plate and form an RBC “button”. If specific antibody is absent, the RBCs will be agglutinated by the virus and form a diffuse film. The titer of the serum or product is the reciprocal of the dilution that completely inhibits agglutination.
HAI is very useful for subtyping influenza virus isolates. A number of factors contribute to the potential variability of the HAI test. Certain serum samples and products may contain nonspecific inhibitors of RBC agglutinins, which may yield false-positive results. A number of procedures have been developed to remove such inhibitors, including adsorption and heat inactivation procedures. Specimens may also contain RBC agglutinins other than specific antibody, and these may contribute to false-negative results. Appropriate preparation and titration of reagents, including RBCs and viral hemagglutinin stocks and suspensions, is critical. In addition, controls for nonspecific agglutination or inhibitors of agglutination must be included in every assay.
Western Blot (or Immunoblot) Assay for Antibody Detection
The immunoblot, or Western blot, assay is a technique for the simultaneous detection of antibodies to various protein antigens of a given virus. The term recombinant immunoblot assay (RIBA) is applicable when the starting protein mixtures are recombinant proteins obtained from prokaryotic or eukaryotic expression systems instead of crude or partially purified virus from infected cells. The method is often used diagnostically as a supplementary or confirmatory test in situations where an initial assay for antibody lacks sufficient specificity or is known to be prone to false-positive results. This is especially important when the test is being used to diagnose an infection of clinical significance such as HIV or HCV infection.
A number of commercial immunoblot kits are available, particularly for viruses such as HIV and HCV; several have regulatory approval for diagnostic use. Alternatively, viral antigen preparations may be produced in-house or purchased, along with other reagents required for the assays. Careful control and/or sourcing of these reagents are critical to ensuring that compliance requirements are maintained.
For selected viral agents, there are generally accepted interpretive standards for the analysis of reactivity or positive results in an immunoblot assay. However, the presence of nonspecific bands may be due to antibody reactivity to cellular protein antigens caused by autoimmune diseases and/or the use of crude virus-infected cell proteins as antigen in the assay. Indeterminate reactions may also occur if only a limited number of specific antibody bands are observed.
Appropriate positive and negative control sera must be included in each assay and reactivity must be scored for both the presence and the intensity of expected protein bands.
Application of the Antibody Detection Methods to Specific Viruses
Human blood-borne pathogens that may be present in infectious form in human donated blood used directly in the production of biological products are a concern because they may present a risk of transmission to others. Testing for virus-specific antibodies in donated blood serves as a screening procedure for the elimination of suspect units. Alternatively, the viruses may represent important agents for which human vaccines have been or are being developed. Thus the ability to detect virus-specific antibodies in an immunized individual or animal model may be important for demonstrating the efficacy of the vaccine. Currently, in the United States, a number of FDA-approved screening or definitive tests may be conducted on donated units of blood for evidence of the presence of agents of infectious diseases, including hepatitis B and C viruses, human immunodeficiency virus, and West Nile virus. In addition, plasma sent for fractionation before production of plasma-derived products is required to be tested for hepatitis A virus (HAV) and human parvovirus B-19.
GLOSSARY
Acceptance Criteria—
Anticipated results, which may be numerical limits, ranges, or other characterization for the tests described. They establish the standards to which a drug substance or drug product should conform in order to be considered acceptable for its intended use.
Adventitious Agent—
Acquired accidental contaminant in a cell line such as viruses and toxins; the agent is often infectious.
Amplicon—
A segment of DNA generated by the PCR process whose sequence is defined by forward and reverse primers.
Antibody—
An infection-fighting protein molecule that binds, neutralizes, and helps destroy foreign microorganisms or toxins. Also known as immunoglobulins, antibodies are produced by the immune system in response to antigens.
Antigen—
Any agent that induces the production of an antibody and reacts specifically with it.
Assay Validation—
A formal, archived demonstration of the analytical performance of an assay that provides justification for use of the assay for an intended purpose and a range of acceptable potency values.
Bioassay—
Analytical method that uses living animals, cells, tissues, or organisms as test subjects.
Biologics—
Products such as antitoxins, antivenins, blood, blood derivatives, immune serums, immunologic diagnostic aids, toxoids, vaccines, and related articles that are produced under license in accordance with the terms of the federal Public Health Service Act (58 Stat. 682) approved July 1, 1944, as amended, have long been known as “biologics.” However, in Table III, Part F, of the Act, the term “biological products” is applied to the group of licensed products as a whole. For Pharmacopeial purposes, the term “biologics” refers to those products that must be licensed under the Act and comply with Food and Drug Regulations—Code of Federal Regulations, Title 21 Parts 600–680, pertaining to federal control of these products (other than certain diagnostic aids), as administered by the Center for Biologics Evaluation and Research or, in the case of the relevant diagnostic aids, by the Center for Devices and Radiological Health of the federal Food and Drug Administration. [Definition from Biologics  1041
1041 , USP–NF vol. 30 (2007), p. 414.]
, USP–NF vol. 30 (2007), p. 414.]
Biotechnology-Derived Product—
Macromolecular article derived from biotechnology processes such as recombinant DNA (rDNA) technology, hybridoma technology, and the like.
Bulk Harvest—
See Unprocessed Bulk Harvest.
Capsid—
The outer protein shell of a virus particle.
Cell Bank—
A defined population of cells, such as an immortalized cell line, grown by a defined process and cryopreserved in a defined process and within a defined passage number range. The assumption is that each vial from a cell bank is comparable and that when thawed and added to a manufacturing vessel (or an analytical assay), it will perform in a consistent way.
Chaotropic—
A reagent that causes molecular structure to be disrupted; in particular, those formed by noncovalent forces such as hydrogen bonding, van der Waals interactions, and the hydrophobic effect.
Complement—
A group of proteins in the blood that work in concert with other immune system proteins and cells (such as antibodies) in attaching foreign substances.
cDNA—
Complementary DNA. Two strands of nucleic acid that can hybridize by specific base pairing between the nucleotides.
Confluency—
Refers to the point when 100% of the surface area of the vessel is covered in cells.
Cryopreservative—
Reagent used to keep a cell alive in deep-frozen condition (usually in liquid nitrogen).
Cytopathic—
Damaging to cells, causing them to exhibit signs of disease or cell death.
ELISA—
Enzyme-linked immunosorbent assay. A biochemical technique used to detect the presence of an antibody or an antigen in a sample.
Endpoint Assay—
An analytical method that measures the amount of accumulated product at the end of the assay.
Epitope—
A molecular region on the surface of an antigen that is recognized by an antibody and can combine with the specific antibody produced by such a response; also called a determinant or an antigenic determinant.
Glycoprotein—
Protein that contains sugar side chains added as a posttranslational process; the presence of sugar side chains often affects activity, antigenicity, and in vivo stability.
Host Cell Tropism—
The range of susceptible cells that a particular microorganism can infect.
ICH—
The International Conference on Harmonization of Technical Requirements for Registration of Pharmaceuticals for Human Use.
Limit of Detection (LOD)—
The lowest concentration of an analyte in a sample that can be detected, not quantitated. It is a limit test that specifies whether or not an analyte is above or below a certain value. It is expressed as a concentration at a specified signal-to-noise ratio, usually 2 or 3.
Mycoplasma—
Parasitic microorganism that infects mammalian cells, possessing some characteristics of both bacteria and viruses. Prokaryotic microorganisms belong to the family Mycoplasmataceae, with no cell walls. They may grow attached or close to cell surfaces in the cytoplasm and subtly change the properties of the cells.
Passage—
An operational procedure used to feed cultured cells, usually by providing fresh medium and dilution of cells in a new culture vessel. The number of such operations is referred to as the passage number. It is not the same as cell generation number, which is strictly related to cell doubling time.
qPCR—
Quantitative polymerase chain reaction. A modification of the polymerase chain reaction used to measure the quantity of DNA, complementary DNA, or ribonucleic acid present in a sample. Like other forms of polymerase chain reaction, the process is used to amplify DNA samples via the enzyme DNA polymerase.
Raw Materials—
All components used to manufacture a drug substance or drug product; regulated by 21 CFR 211.
RT-PCR—
Reverse transcriptase polymerase chain reaction. A variation of the PCR technique in which cDNA is made from RNA via reverse transcription. The cDNA is then amplified using standard PCR protocols.
Serotype—
The kind of microorganism as characterized by testing for recognizable antigens on the surface of cells of the microorganism.
Spiking—
Adding a known amount of analyte from a laboratory standard acting as a tracer to check a method for recovery or accuracy.
Syncytium—
A multinucleated mass of cytoplasm that is not separated into individual cells.
System Suitability—
The checking of a system to ensure system performance before or during the analysis of unknowns. Parameters such as plate count, tailing factors, resolution, and reproducibility are determined and compared against the specifications set for the method. These parameters are measured during the analysis of a system suitability sample, which is a mixture of main components and expected by-products.
TCID50—
50% tissue culture infective dose. The level of dilution of a virus at which half of a series of laboratory wells contain active, growing virus.
Unprocessed Bulk Harvest—
The pooled harvests of cell culture fluids that constitute a homogeneous mixture for manufacture into a unique lot of product.
APPENDIX: RELEVANT REGULATORY REFERENCES
- CBER (Center for Biologics Evaluation and Research), U.S. Food and Drug Administration. Points to Consider in the Characterization of Cell Lines Used to Produce Biologicals. 1993.
- CBER (Center for Biologics Evaluation and Research), U.S. Food and Drug Administration. Letter to manufacturers of biological products. 2000.
- 9 CFR 113.53.
- 9 CFR 113.47.
- 9 CFR 113.52.
- 21 CFR 211.
- 21 CFR 600.3.
- Federal Food, Drug, and Cosmetic Act (FD&C Act), sections 201 (g) and (h).
- Guidance on Virus Validation Studies (CPMP/BWP/268/95 EMEA, Feb. 1996).
- International Conference on Harmonization of Technical Requirements for Registration of Pharmaceuticals for Human Use (ICH). Q5A (R1): Viral Safety Evaluation of Biotechnology Products Derived from Cell Lines of Human or Animal Origin. 1997.
- Public Health Service Act, section 351 (i).
-
Viral Safety Evaluation of Biotechnology Products Derived from Cell Lines of Human or Animal Origin
 1050
1050 . USP–NF vol. 30 (2007), pp. 491–500.
. USP–NF vol. 30 (2007), pp. 491–500.
Auxiliary Information—
Please check for your question in the FAQs before contacting USP.
| Topic/Question | Contact | Expert Committee |
|---|---|---|
| General Chapter | Fouad Atouf, Ph.D.
Senior Scientific Liaison 1-301-816-8365 |
(GCBA2010) General Chapters - Biological Analysis |
USP35–NF30 Page 919
Pharmacopeial Forum: Volume No. 34(2) Page 391